Team Brain Genomics
The Brain Genomics research group investigates the influence of genomic variability on inter-individual differences in the structure and function of the human brain.

The research group „Genomic Imaging“ studies the influence of genetic factors on structure and function of the human brain.
For this purpose, we combine cutting edge molecular genetic techniques, structural and functional brain imaging as well as neuropsychological data. Our research aims at understanding the neurobiological basis of phenotypic variability of the healthy and diseased brain. Further, genetic, transcriptomic and epigenetic data support the formation of a multiscale and multimodal digital atlas of the human brain. This atlas integrates different data levels and will be available to users.
Imaging techniques have revealed remarkable structural and functional variability in the brain. Heritability estimates provide insights into the quantitative contribution that genetic factors have on the observed variability. To systematically search for common genetic variation that influences that variability, we employ genome-wide association studies (GWAS). For these studies, it is crucial to have large, well-characterized samples at hands, and we use the 1000Brains,as well as samples available through collaborations. We are also involved in the international ENIGMA consortium.
Recently, the systematic study of the impact of rare genetic variants has become increasingly important, we employ whole exome and whole genome sequencing (next generation sequencing – NGS) for their identification.
The biostatistics and bioinformatics analysis of the data is particularly challenging because most phenotypes are influenced by a complex interplay of many genetic factors. Apart from that, environmental factors also play a role. There is a growing need for the application of statistical approaches that can handle this enormous complexity, such as deep learning. This requires high performance computing (HPC) which is available at the Jülich Supercomputing Center (JSC).
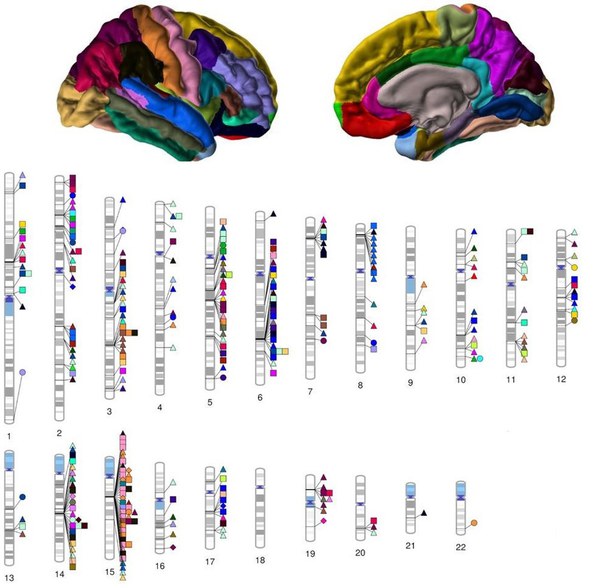
Genetic studies in large cohorts of patients with neuropsychiatric and neurological diseases identify an increasing number of disease-associated genetic variants. We study the influence of such variants on the phenotypic variability observed in healthy individuals. For these studies, the availability of large, well-characterized cohorts of individuals derived from the normal population, such as the above mentioned 1000Brains study, is of great value.
Regional differences of gene expression and epigenetic patterns in the human brain. In the EU flagship Human Brain Project we study regional differences in gene expression and DNA methylation in post-mortem brain samples. These data are incorporated in the Human Brain Atlas. Knowledge of such regional differences provides the basis for the recognition of gene groups and biological pathways that contribute to the functional differences. In this context, we have developed the new bioinformatic tool JuGEx in collaboration with the research group Architecture and Brain Function.