Aptamers are short DNA or RNA oligonucleotides with the capability to selectively bind specific target molecules and are considered as synthetic antibodies. They are usually obtained by a selection from a large random sequence library accordingly to their binding capabilities to the target molecule via a SELEX process (Systematic Evolution of Ligands by Exponential). The aptamer molecules can be utilized as receptors for biosensors similar to normal antibodies, however with the advantage of being more robust, affordable, and relatively tiny. The small size is of great advantage for the development of electrochemical biosensors since they are more confined in the vicinity of the electrode and render high receptor densities thus allowing to measure high signals. We develop novel biosensors concepts that are based on changes of the solid/liquid interface impedance caused by aptamer/target binding events. Those changes of the sensor surface composition are detected via different transducer such as liquid gate FETs, organic electrochemical transistors (OECTs), micro-fabricated microelectrodes, and conventional electrochemical three-electrode setups.
Electrochemical Aptasensors

Amperometric aptasensors have become very popular recently due to their capability to detect minute amounts of analyte molecules. These sensors typically exploit conformational changes of the aptamer induced by the binding of target molecules. If a redox probe is attached to the distal end of the surface tethered aptamer receptor, then the conformational rearrangement evokes a variation of the charge transfer resistant between redox probes and electrode. The resulting redox current can be recorded as sensor signal. We are developing new schemes for signal amplification by means of redox cycling, electrochemical rectification, and nanomaterials in order to improve signal reliability, to the extend dynamic range of detection, and to lower the detection limit.
Our target systems are:
- Malaria biomarkers: Plasmodium falciparum lactate dehydrogenase (PfLDH), Plasmodium vivax lactate dehydrogenase (PvLDH), and P. falciparum histidine-rich protein 2 (HRP-2);
- COVID-19 biomarkers: Spike glycoprotein of SARS-CoV-2 virus;
- Alzheimer disease biomarkers: amyloid beta oligomers (AbO);
- Neurotransmitters: glutamate, dopamine (DA), ATP, serotonin;
- Diabetes biomarkers: insulin, glycated human serum albumin;
- Tumor Marker: VEGF
Aptamer adaptation
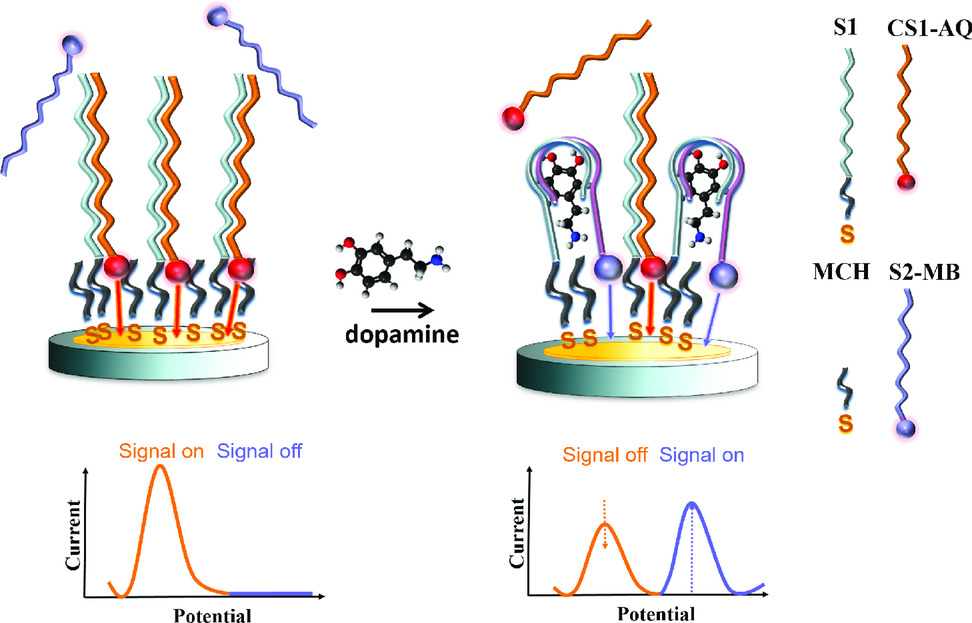
Aptamers offer great flexibility in terms of structure variability with high affinity and specificity for their analyte molecules. Furthermore, the structure and thus functionality can be synthetically modified by changing the aptamer sequence or by attaching diverse functional groups. Alternatively, the aptamer can be split into two fragments to decouple the binding process from the detection. Those two ssDNA fragments can form associated complexes with their target building a sandwich-like structure. In our work, a whole dopamine aptamer was divided into two parts. One of those aptamer fragments was modified with a thiol binding group (S1) used as capture stand for surface immobilization. The second part of the aptamer carried a redox probe (S2-MB) indicating the binding of the target molecule similar to the detection antibody in a binding immunosorbent assay (for instance ELISA). Aptamer splitting improved the signal to noise ratio of the sensor and reduced the detection limit. Furthermore, additional redox probes can be introduced to the linked ssDNA sequences (CS1-AQ) complementary to the capture strand tethered on the sensor surface. Those complementary strands provided an amperometric signal which was proportional to the number of immobilized capture strands, thus permitting an independent evaluation of the sensor status before testing.
Electrochemical Logic Gates
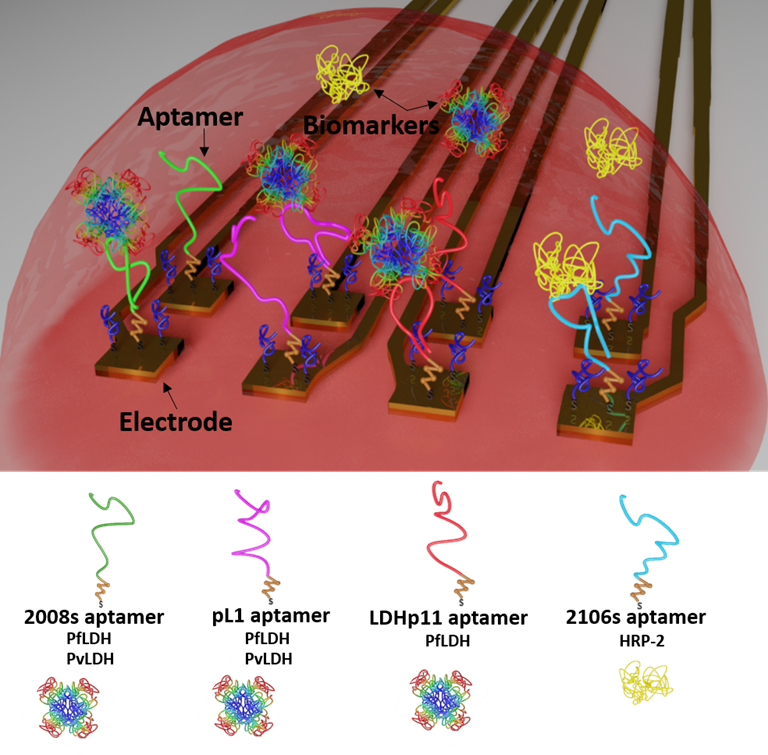
Electrochemical logic gates can be used to combine serveral sensor signals to improve the reliability of sensor outputs by means of signal amplification, multi-input integration, failure checking, and many more. We developed a multi-target malaria biosensor system, to distinguish between the two main malaria parasites, P. falciparum and P. vivax. Specific Plasmodium detection is achieved by immobilizing four distinct and specific aptamers on individual electrodes of a flexible multielectrode array (flexMEA) chip. Each electrode represents an independent aptasensor that specifically or selectively detect a biomarker from the malaria parasites and its signal is recorded separately. When exposing the sensor array to a blood sample, the combination of different output signals from the different aptasensors can be operated as logic gates for specific parasite detection. The importance of this distinctive identification allows guiding a correct antimalarial treatment to avoid the drug resistance problem.
In another example, we have demonstated a three level cascade logic gate that converts a relatively complex set of information into a simple YES or NO diagnosis.
Dr. Dirk Mayer
Tel.: +49-2461-61-4023
e-mail: dirk.mayer@fz-juelich.de
PUBLICATIONS:
Ratiometric electrochemical aptasensor with strand displacement for insulin detection in blood samples. Zhou, L., et al. Analytica Chimica Acta (2024): 342823.
Flexible multielectrode arrays based electrochemical aptasensor for glycated human serum albumin detection. Zhou, L., et al. Sensors and Actuators B:Chemical 386 (2023): 133730.
Delineating charge and capacitance transduction in system-integrated graphene-based BioFETs used as aptasensors for malaria detection. Figueroa-Miranda, Gabriela, et al. Biosensors and Bioelectronics 208 (2022) 114219.
Highly selective and sensitive detection of glutamate by an electrochemical aptasensor. Wu, Changtong, et al. Analytical and bioanalytical chemistry 414 (2022) 1609-1622.
Multi-target electrochemical malaria aptasensor on flexible multielectrode arrays for detection in malaria parasite blood samples. Figueroa-Miranda, Gabriela, et al. Sensors and Actuators B: Chemical 349 (2021) 130812.
Ultrasensitive antibody-aptamer plasmonic biosensor for malaria biomarker detection in whole blood. Minopoli, Antonio, et al. Nature communications 11 (2020) 1-10.
Polyethylene glycol-mediated blocking and monolayer morphology of an electrochemical aptasensor for malaria biomarker detection in human serum. Figueroa-Miranda, Gabriela, et al. Bioelectrochemistry 136 (2020) 107589.
A novel ratiometric electrochemical biosensor based on a split aptamer for the detection of dopamine with logic gate operations. Guo, Ting, et al. Physica Status Solidi (a) 217 (2020) 1900924.
Electrochemical dual-aptamer biosensors based on nanostructured multielectrode arrays for the detection of neuronal biomarkers. Zhang, Yuting, et al. Nanoscale 12 (2020) 16501-16513.
Amperometric aptasensor for amyloid-β oligomer detection by optimized stem-loop structures with an adjustable detection range. Zhang, Yuting, et al. ACS sensors 4 (2019) 3042-3050.
Monitoring amyloid-β proteins aggregation based on label-free aptasensor. Zhang, Yuting, et al. Sensors and Actuators B: Chemical 288 (2019) 535-542.
Aptamer-based electrochemical biosensor for highly sensitive and selective malaria detection with adjustable dynamic response range and reusability. Figueroa-Miranda, Gabriela, et al. Sensors and Actuators B: Chemical 255 (2018) 235-243.
Biosensing near the neutrality point of graphene. Fu, Wangyang, et al. Science advances 3 (2017) e1701247.
Electrochemically triggered aptamer immobilization via click reaction for vascular endothelial growth factor detection. L. Feng et al, Engineering in Life Sciences. 16, (2016) 550-559
Multi‐Level Logic Gate Operation Based on Amplified Aptasensor Performance. L. Feng et al, Angew. Chem. Int. Ed., 54, (2015) 7693-7697.
Electrochemical current rectification–a novel signal amplification strategy for highly sensitive and selective aptamer-based biosensor. L. Feng et al, Biosensors and bioelectronics, 66, (2015) 62-68.
