OneProt: Towards Multi-Modal Protein Foundation Models
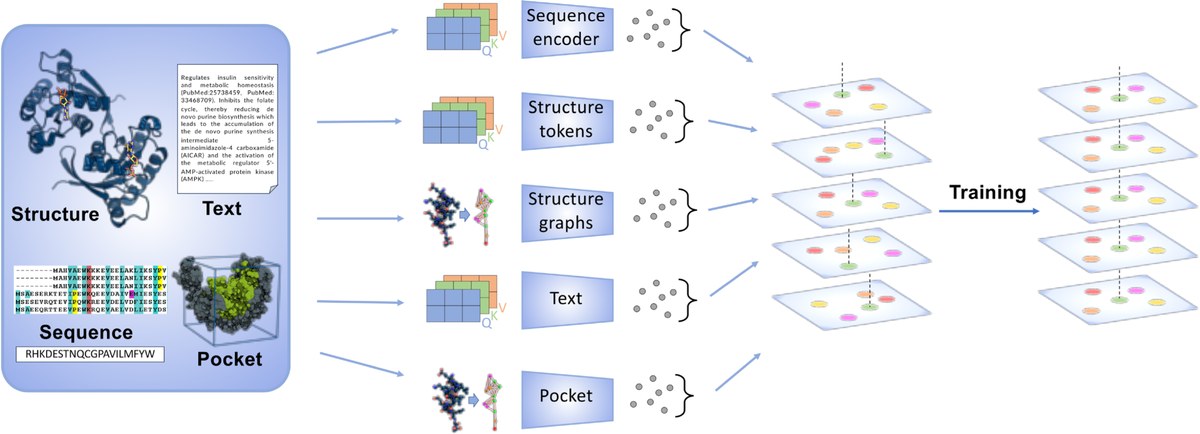
A new study conducted by Helmholtz AI researchers and published in PLOS Computational Biology, presents OneProt, a new and flexible AI system designed to help scientists better understand proteins. OneProt brings many kinds of information about proteins together—such as their 3D structures, amino acid sequences, written descriptions, and binding site details—by using the ImageBind framework. This allows the system to align different data types efficiently, even when they do not match perfectly.
By combining Graph Neural Networks with transformer models and leveraging the computing power of JUWELS Booster, OneProt performs especially well on tasks like predicting enzyme function and analyzing protein binding sites. It can also share information across different data types, making it easier to spot similarities between proteins.
The new AI system offers two major advantages: it can incorporate new kinds of data during pre-training, and it can be fine-tuned easily using just a small neural network layer. The study also shows that using multiple types of data reduces the need for very large training datasets while still achieving strong performance. Additionally, through a detailed ablation study, the importance of the binding site encoder was highlighted—an innovation not seen in similar models. Overall, OneProt marks a meaningful step forward in multi-modal protein modeling, with strong potential for advancing drug discovery and protein engineering.
The next generation project following OneProt, OneProtGPT, is a part of the Gauss AI Compute Competition and is currently running on JUPITER. It is a powerful AI model that connects protein data with large language models, enabling a deeper understanding of proteins across multiple types of information and allowing scientists to generate consistent and enriched descriptions of proteins from diverse inputs. Its applications include designing new proteins and optimizing industrial enzymes, making it a potentially valuable tool for both research and industry.
The article is available here (open access): OneProt: Towards multi-modal protein foundation models via latent space alignment of sequence, structure, binding sites and text encoders. Flöge K, Udayakumar S, Sommer J, Piraud M, Kesselheim S, et al. (2025). PLOS Computational Biology 21(11): e1013679. https://doi.org/10.1371/journal.pcbi.1013679
Contact: Alina Bazarova
