Membranes
Wrapping of Nanoparticles by Lipid-Bilayer Membranes
Membrane budding and particle uptake is important for the communication of cells with their environment, e.g. for endocytosis, phagocytosis, and parasite or virus entry. Also, various potential applications of nanoparticles in complex materials require a better understanding of their cellular toxicity. Furthermore, nanoparticles can be used for targeted drug delivery, for cancer therapy, and as membrane-makers for biomedical studies.
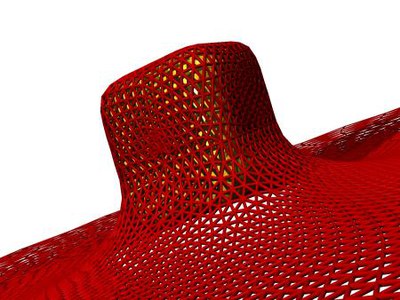

We study the passive endocytosis of nanoparticles between ten and a few hundred nanometers in size that are wrapped in lipid bilayer membranes. Upon wrapping, the membrane deformation energy increases while the adhesion energy due to the attractive interaction between particle and membrane decreases. Numerical calculations with triangulated surfaces allow the calculation of deformation energies for various particle shapes.
We characterise nanoparticle wrapping analogously to thermodynamic phase transitions. While partially-wrapped states are found only in membranes with tension in the case of spherical nanoparticles, cube-like, rod-like, and ellipsoidal nanoparticles show shallow and deep partially-wrapped states also without tension. Here, shape matters! Not only the aspect ratio, but also the local curvature distribution on the nanoparticle surface controls binding and wrapping. This is the first crucial step for the interaction of nanoparticles with biological cells.
Fluctuating shells under pressure
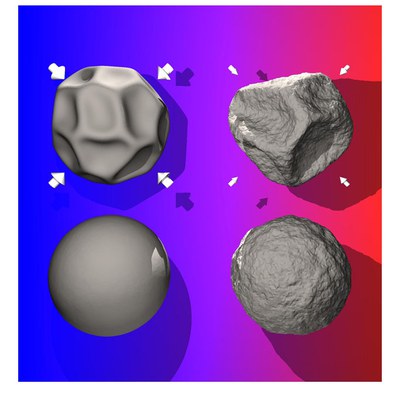

A thin spherical shell can be considered as an elastic membrane with shear modulus, bending rigidity and a non-zero curvature. On a micrometer and submicrometer scale, examples of these shells range from synthetic hollow polyelectrolyte capsules with important technological applications, through to biologically relevant systems such as red blood cells and spherical virus capsids. In the absence of thermal fluctuations, the stability of elastic shells against external forces such as a uniform pressure field or a point-like indentation depends on the ratio between the size of the shell and the thickness of the wall. As in flat membranes, thermal fluctuations are expected to influence the mechanical response of a deformed shell by the renormalization of elastic constants.
However, the fluctuations of thin shells are qualitatively different from those in flat elastic membranes due to the coupling of in-plane stretching modes and bending modes by the curvature.We study the deformations of these shells using Monte Carlo computer simulations and perturbation theory, including the effects of curvature and external pressure. We show that thermal fluctuations reduce the critical buckling pressure and soften the mechanical response on point-like indentations.
Budding Induced by Conical Inclusions
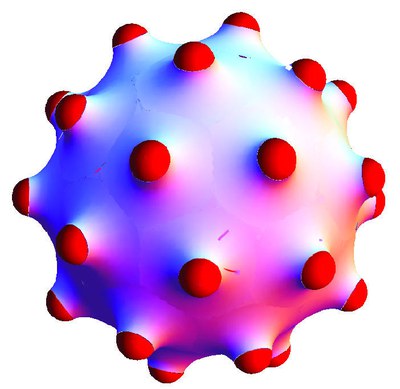
Budding and vesiculation of lipid bilayers is the first step for material transport in biological cells. Bud formation can be induced by curved inclusions, such as asymmetric proteins or viruses that are partially adhered to the bilayer.
Whereas curved inclusions in a planar membrane experience a repulsive membrane-mediated interaction, we find an effective attraction for inclusions on a bud. This attraction is caused by the screened repulsion of inclusions on a curved membrane. Using a model that contains bending energy only, we find an optimal bud radius and therefore a spontaneous curvature c0 for a given density of inclusions, corresponding to catenoid-like membrane-deformation patches around the inclusions. For budding from a mother vesicle, bending energy alone generates a line of degenerated ground states for the bud radius as a function of number of inclusions per bud. Translational entropy of the inclusions lifts that degeneracy and free-energy minimization then allows a unique prediction of bud radii.
Crumpling of thin elastic sheets
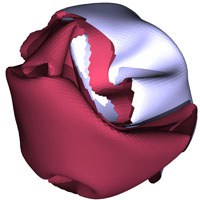

The deformation of thin elastic sheets is a fundamental problem with many practical applications to different physical and biological systems. Among these systems are macroscopic materials extending from thin steel plates via thin rubber films to paper sheets, mesoscopic materials like clay platelets, the membrane of biological cells and giant vesicles, but also microscopic materials like virus particles and carbon nano-tubes.
Using computer simulations we investigated the effect of self-avoidance on the shapes, mechanical properties and fold length distributions of crumpled elastic sheets.
Buckling of virus capsids
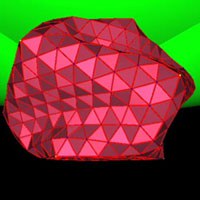

The genome of a virus is contained in a protective cage known as the viral capsid. A viral capsid has a remarkably regular structure and is built up from a fixed number of copies of a single or a few kinds of capsid protein. Its geometry is that of an icosahedron or a helical cylinder, but more complex structures also exist. We investigate the mechanical properties of icosahedral virus shells by computer simulations. We predict the elastic response for small deformations, and the buckling transitions at large deformations which depend both strongly on the number of elementary building blocks, the shear and bending elasticity of the shell and the confining geometry.
Polymer-decorated membranes
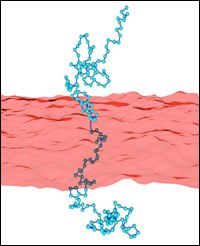
Polymer-membrane interactions are relevant in many complex systems, ranging from membranes of biological cells to microemulsions. For example, polymer boosting in ternary microemulsions enhances the oil-water mixing efficiency of the surfactant. Polymers anchored to membranes suppress fluctuations of the membrane conformation, i. e. polymer addition effectively stiffens the membrane. Using Monte-Carlo simulations, we determine effective curvature-elastic constants for polymer-decorated membranes. In particular, we investigate the effect of self-avoidance within a single polymer chain. This has been extended to investigate other polymer architectures like star polymers. With increasing arm number of anchored stars, the bending rigidity is found to increase strongly, while the saddle-splay modulus remains constant.
Budding of crystalline domains in fluid membranes

In biological cell, the formation of small vesicles at the plasma membrane, a process which is called endocytosis, is controlled by the adsorption of clathrin proteins. These molecules adsorb on the membrane to form a domain of a regular, hexagonal network. Thus, the formation of vesicles from such a domain corresponds physically to the budding of crystalline domains embedded in fluid membranes. Here, budding is driven by the line tension of the domain boundary, and by the spontaneous curvature c0 of the domain. We employ Monte Carlo simulations of dynamically triangulated surfaces and scaling arguments to study the budding process. In a crystalline phase, the formation of a spherical shell requires in appearence of lattice defects. In a triangular lattice, which in the flat phase has only 6-fold coordinated vertices, these are 5-fold and 7-fold coordinated "disclinations" (see figure). We find that the defects are generated at the boundary, and then diffuse into the interior. In contrast to previous theoretical predictions, the budding is found to occur at a value of the spontaneous curvature c0, which is a decreasing function of the domain size.
Advanced Flicker Spectroscopy of Fluid Vesicles
Shapes and fluctuations of fluid membranes are controlled by their curvature elasticity, which is characterized by two elastic constants, the bending rigidity and the spontaneous curvature. In turn, the spectrum of membrane fluctuations provides information about the values of the elastic constants. The analysis of membrane fluctuations - which is called flicker spectroscopy - has been limited so far to quasi-spherical vesicles, which allows only to extract the bending rigidity, but is insensitive to the spontaneous curvature. We have developed a a new method, advanced flicker spectroscopy of giant, non-spherical vesicles, which makes it possible to measure both parameters simultaneously. Our analysis is based on the generation of a large set of reference data from Monte Carlo simulations of randomly triangulated surfaces. The method has been applied to thermal trajectories of vesicle shapes and to the elastic response of zwitterionic membranes to trans-membrane pH gradients. The new technique makes it possible to easily characterize membrane curvature as a function of environmental conditions.
Water Molecules and Hydrogen-Bonded Networks in Bacteriorhodopsin -
Molecular Dynamics Simulation Completes Crystal Structure
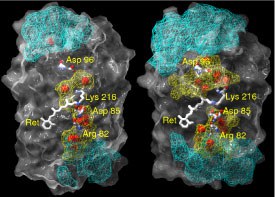
The spatial distribution of water molecules and their corresponding hydrogen-bonded network inside bacteriorhodpsin (bR) in its ground state conformation were investigated by molecular dynamics simulation. The protein was embedded in a fully hydrated lipid bilayer membrane. A much higher average number of internal water molecules (44) than observed in crystal structures (18) was found in our simulation. This discrepancy is due to the high mobility of these water molecules between different positions. On average we found 20 water molecules trapped and 24 diffusive inside bR. The time of exchanging the whole ensemble of diffusive internal water molecules is of the order of 200 ps. The average residence time of a diffusive water molecule inside the protein is about 52 ps.
We have characterized the hydrogen-bonded networks inside the protein based on the Grotthuss model as a geometrical construct. It is shown that the length of one hydrogen-bonded path connecting a charged residue to the cytoplasmic surface consists in average of 5 bonds. The average lifetime of an uninterrupted hydrogen-bonded pathway is of the order of 0.045 ps. The average spatial distribution of hydrogen-bonds between the cytoplasmic surface and Asp96 suggests a kind of funnel directed towards this residue. This study provides a new basis for the transduction of protons via highly fluctuating hydrogen-bonded networks and thus introduces a new element to the mechanism of proton pumping. The ground state was chosen in this study to demonstrate that even the structure with the lowest amount of water molecules, as seen in the crystal structure, contains a dense and highly fluctuating hydrogen-bonded network in an almost hydrophobic part of bacteriorhodopsin (2004).
