Protein dynamics
Large scale motions of globular domains are often crucial for the activity of multidomain proteins. We have employed single-molecule Förster resonance energy transfer (smFRET) for the characterization of large scale domain motions.
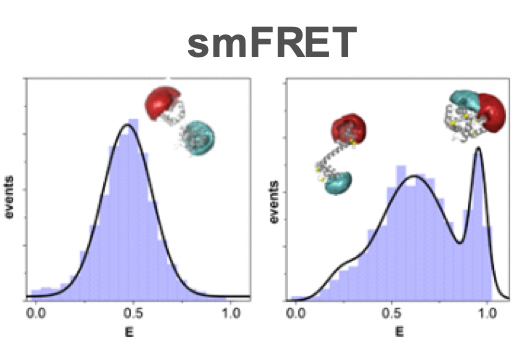
A successful application of the FRET spectroscopy is given by the analysis of ligand induced conformational changes in so-called sensor proteins. These proteins can be employed to determine the ligand concentration, for example for glucose, calcium, or special amino acids. By this, sensor proteins can be utilized to characterize the chemical composition within the cytosol. Many of these sensor proteins make use of the venus flytrap principle: The ligand induces a bending and a swiveling twist motion about the hinge with an open conformation in the ligand-free state and a close conformation in the ligand-bound state. To measure this conformational change, variants of a cyan fluorescent protein (FP) and a yellow FP (acceptor), are fused to the amino- and carboxyl-termini of the sensor protein, respectively. The design of this kind of sensors is still a challenge and requires the application of sophisticated methods.
Related Publications
- Reinartz I., Sarter M., Otten J., Höfig H., Pohl M., Schug A., Stadler A.M., Fitter J. (2021) Structural Analysis of a Genetically Encoded FRET Biosensor by SAXS and MD Simulations. Sensors, 16;21(12):4144
- Höfig H., Otten J., Steffen V., Pohl M., Boersma A.J., Fitter J. (2018) Genetically Encoded Förster Resonance Energy Transfer-Based Biosensors Studied on the Single-Molecule Level. ACS Sens., 3, 1462-1470
- Gabba M., Poblete S., Rosenkranz T., Katranidis A., Kempe D., Züchner T., Winkler R.G., Gompper G., Fitter J. (2014) Conformational State Distribution and Catalytically Relevant Dynamics of a Hinge-Bending Enzyme Studied by Single-Molecule FRET and a Coarse-Grained Simulation. Biophys. J., 107, 1913-1923
- Inoue R., Biehl R., Rosenkanz T., Fitter J., Monkenbusch M., Radelescu A., Farago B., Richter D. (2010) Large domain fluctuations on 50 ns timescale enable catalytic function in phosphoglycerate kinase. Biophys. J. 99, 2309-2317
